Foundation Models for Spatial Intelligence
Spatial correspondence estimation as infrastructure for healthcare AI.
I develop foundation models for spatial alignment that generalize across anatomies, imaging modalities, and clinical tasks. Instead of training a new model for each dataset, our goal is to learn a general capability for estimating spatial correspondence from diverse data, enabling out-of-the-box alignment on new problems.
The resulting deformation fields, however, are more than a registration output. As a structured spatial representation, they encode anatomical variation, motion, and structural change — making them a natural substrate for downstream tasks such as biomechanical model fitting, physics-based simulation, surgical navigation, and longitudinal analysis.
I see this line of work as a step toward spatial foundation models: general-purpose infrastructure for spatial intelligence in healthcare, where labeled data and task-specific supervision are rarely available.
Registration Foundation Models
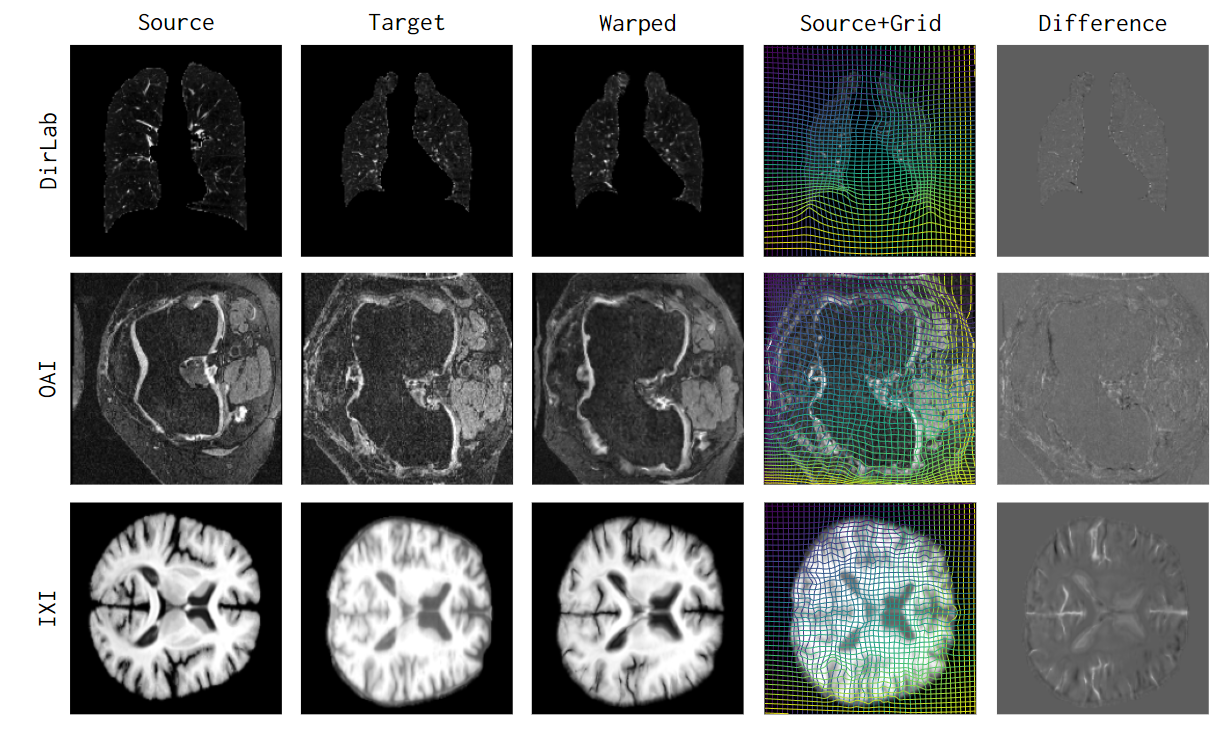
uniGradICON (Tian et al., 2024) is a generalizable registration foundation model supporting diverse anatomies (brain, lung, abdomen, knee) and imaging modalities without retraining.
multiGradICON (Demir et al., 2024) extends this framework to multimodal registration.
Reliable Deployment
For foundation models to be used in clinical and scientific settings, it is important to quantify when the model may fail.
Test-time uncertainty estimation (Tian et al., 2026) via transformation equivariance enables plug-and-play uncertainty estimation, turning pre-trained registration networks into risk-aware tools without modifying the original models.
References
2026
- arXiv
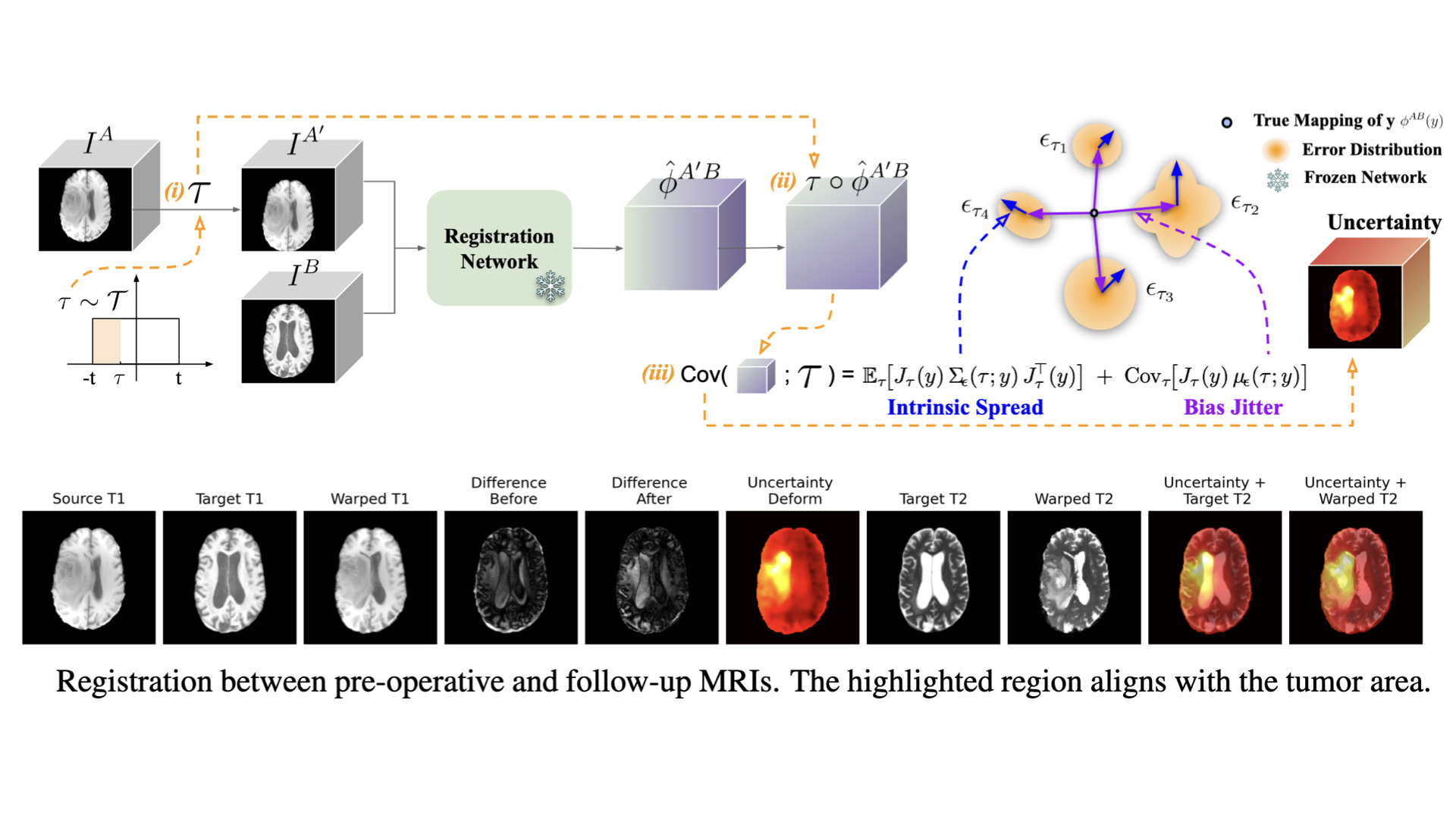 Test-time Uncertainty Estimation for Medical Image Registration via Transformation EquivariancearXiv preprint arXiv:2509.23355, 2026
Test-time Uncertainty Estimation for Medical Image Registration via Transformation EquivariancearXiv preprint arXiv:2509.23355, 2026
2024
- WBIRmultiGradICON: A Foundation Model for Multimodal Medical Image RegistrationIn Workshop on Biomedical Image Registration – WBIR 2024, 2024Oral Presentation