Physics-based Simulation
Physics-based simulation of anatomy, imaging, and deformation for healthcare applications.
I develop physics-based simulation methods to model anatomical deformation and medical image formation. These simulations enable data generation, algorithm development, and validation in applications where real data are limited or difficult to acquire, such as surgical navigation and neuropathology–neuroimaging joint analysis. More broadly, this line of work moves toward digital twins of anatomy and imaging processes, enabling simulation-driven learning and model-based reasoning in healthcare.
Surgery Navigation
I develop physics-based imaging simulation and differentiable rendering methods to generate paired data for cross-modality and cross-dimensional registration problems in surgical navigation.
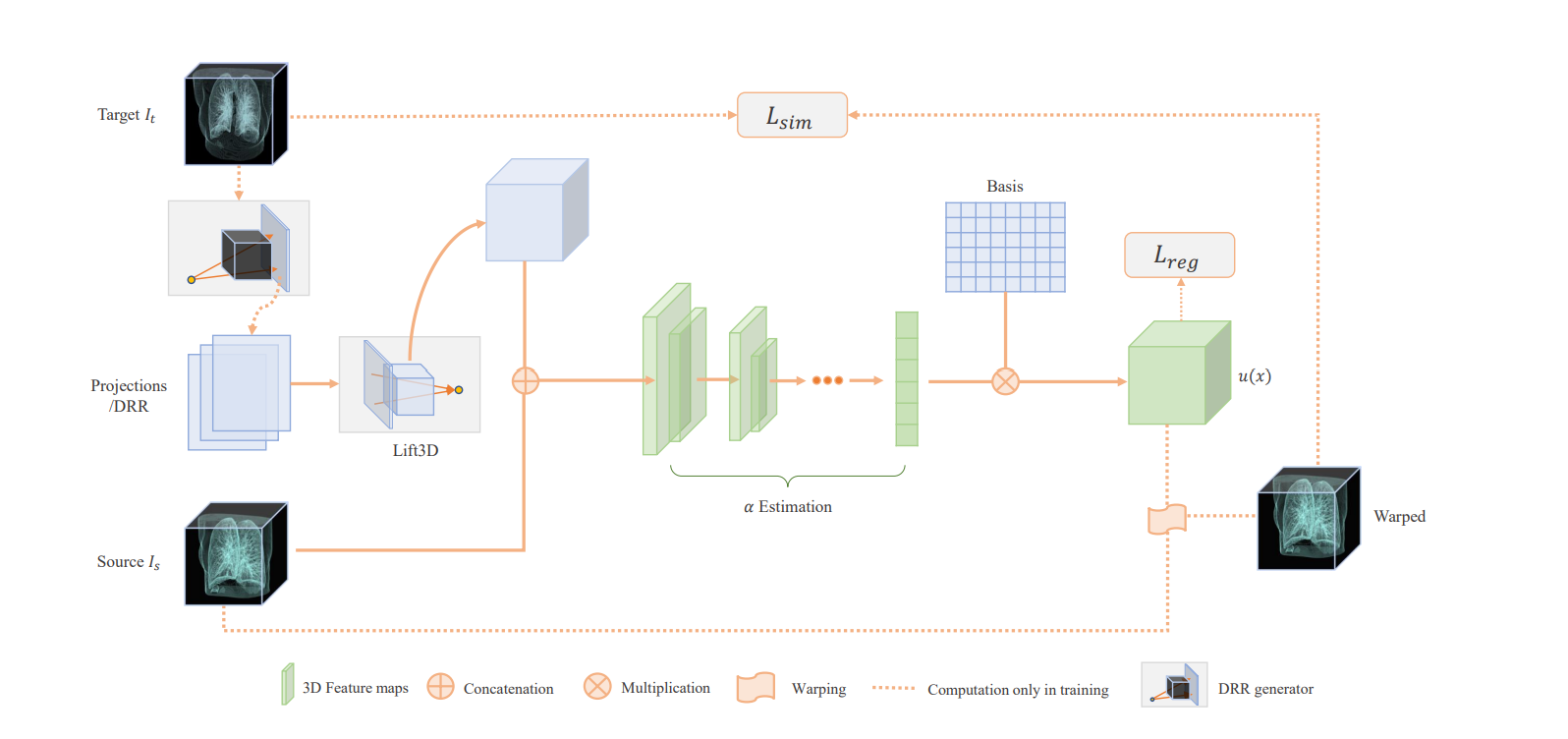
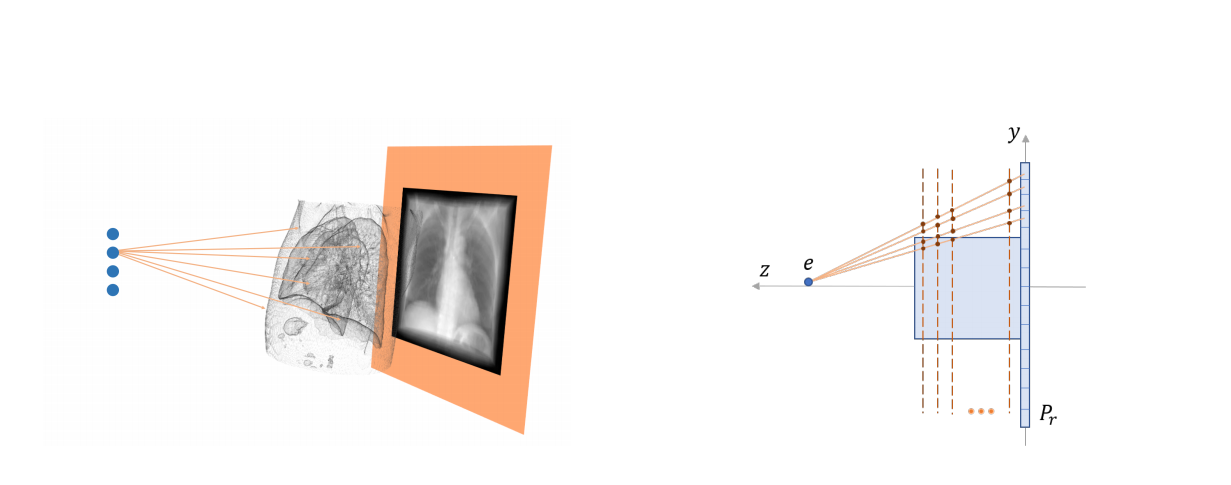
I build radiograph projection simulation from CT images via differentiable rendering, which enables fluid registration (Tian et al., 2020) between preoperative lung CT and intraoperative stationary chest tomosynthesis images during respiration. We later developed LiftReg (Tian et al., 2022), a real-time registration network that aligns 3D CT with intraoperative limited-angle tomosynthesis images using simulated paired CT/projection datasets.
Neuropathology-neuroimaging Joint Analysis
I develop simulation and domain randomization techniques to enable spatial reconstruction and alignment in problems where paired data cannot be directly acquired.
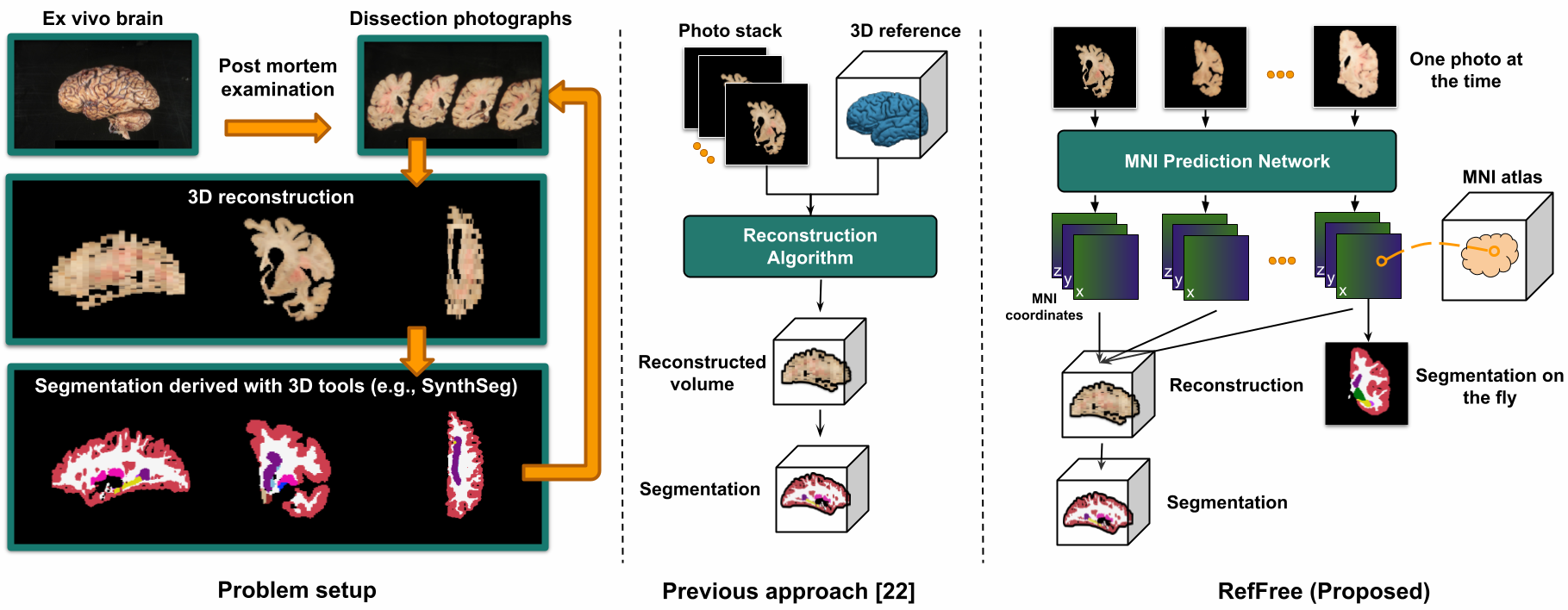
Reference-free 3D reconstruction (Tian et al., 2025) of brain dissection photographs enables spatial localization of neuropathology (e.g., Alzheimer’s disease, stroke) by aligning reconstructed volumes with in vivo MRI. Since acquiring large-scale ex vivo brain dissection photographs relies on postmortem donations and is inherently limited, we use domain randomization to train the correspondence prediction network purely from in vivo MRI data.